Why GenPipes?
Genomic sequencing has become an indispensable tool for modern bioinformatics researchers in their quest to understand biological processes. Next generation sequencing (NGS) is not only complex but is also compute and data intensive. It requires efficient handling of high performance computing infrastructure, managing scalability, flexibility and capability to handle massive amounts of genome reference data while managing intermediate results and inter-dependency between several serial and parallel analysis processes.
GenPipes is a python based bioinformatics tool comprising of several pre-built pipelines that addresses genomic analysis needs for bioinformatics researchers. It is available for public use as open source software. GenPipes was developed at the Canadian Centre for Computational Genomics (C3G). It offers a wide array of genomic sequencing pipelines including RNA-Seq, ChIP-Seq, Whole Genome Sequencing (WGS), Exome sequencing, Bisulfite sequencing, Hi-C, capture Hi-C, metagenomics and SARS-CoV-2 genome sequencing pipeline.
GenPipes Features
Multiple Schedulers - GenPipes is optimized for HPC processing. Currently, it supports 4 schedulers - Slurm, PBS/Torque, Batch, Daemon.
Optimal Job Execution Time - GenPipes minimizes overall job analysis time by job dependency model that leverages job parallelism. This enables jobs to become active and executed as soon as the dependencies are met.
Smart Relaunching of Jobs - Through smart tracking of job progress, GenPipes can determine which jobs failed and at which steps. This helps to relaunch the jobs at the exact point in time, just before the last failure, automatically.
Parameter Encapsulation - GenPipes is a flexible framework that allows user adjustments. It implements a superposed configuration system to reduce the time required to set-up or modify parameters needed during the analysis.
Diverse Inputs - GenPipes is flexible in terms of multiple choice of input files for the analysis. It allows users to skip specific steps in the pipeline if they consider them unnecessary.
Customizable Workflows - GenPipes limits wastage of expensive HPC resources and time as it allows customizable steps in different pipelines enabling users to plug and play with customized pipeline steps.
Key Differentiators
GenPipes is different from other analysis platforms, workbenches and workflow management systems (WMS) in terms of the following capabilities:
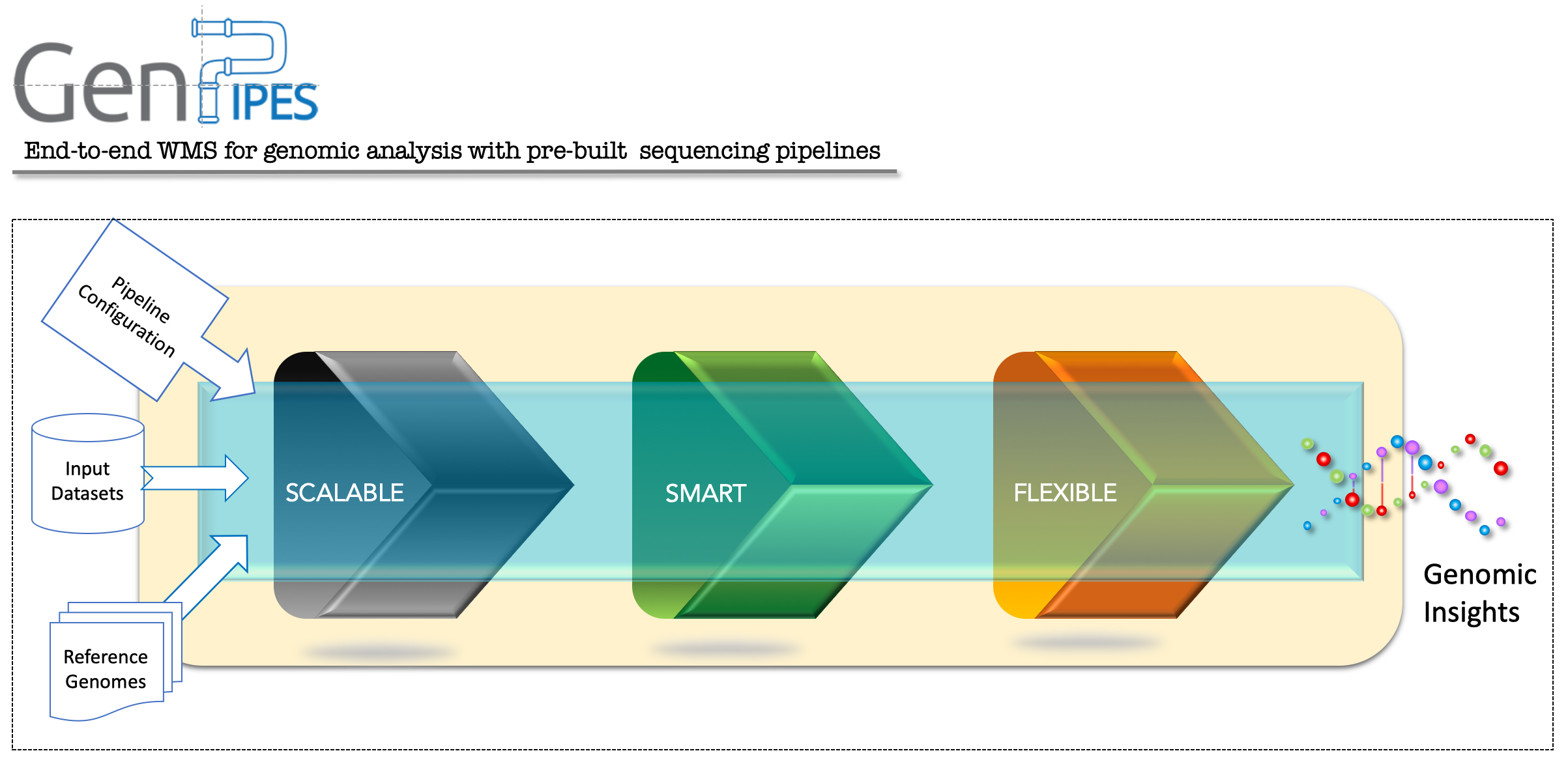
Flexibility: Easy to modify and configure, multiple type of deployments available – local (containerized), cloud (GCP) and hosted on Compute Canada data centre, support for multiple job schedulers
Scalability: Optimized for large scale data analysis, simple to scale up or down in terms of processing and data access needs.
Built-in Pipelines: Pre-built, tested on multiple computing infrastructures, robust industry standard benchmark driven and production quality genomic analysis pipelines for various bioinformatics analyses.
Comparing GenPipes with other NGS Solutions
GenPipes’ strength lies in its robust WMS that comes with one of the most diverse selection of analysis pipelines that have been thoroughly tested. The pipelines in the framework cover a wide range of sequencing applications. The pipelines are end-to-end workflows running complete bioinformatics analyses. While many available pipelines conclude with a bam file or run limited post-bam analysis steps, the pipelines included in GenPipes are extensive, often having as many as 40 different steps that cover a wide range of post-bam processing.
For a tabular comparison of available solutions for NGS see here.
GenPipes is compatible with HPC computing, as well as cloud computing, and includes a workflow manager that can be adapted to new systems. GenPipes also provides job status tracking through JSON files that can then be displayed on a web portal (an official portal for GenPipes will be released soon). GenPipes’ available pipelines facilitate bioinformatics processing, while the framework makes it flexible for modifications and new implementations.
GenPipes developers offer continuous support through a Google forum page and a help desk email address (pipelines@computationalgenomics.ca). Since the release of version 2.0.0 in 2014, a community of users has run GenPipes to conduct approximately 3,000 analyses processing ∼100,000 samples.
To learn more about what GenPipes is and how it works, refer to the Getting Started Guide.